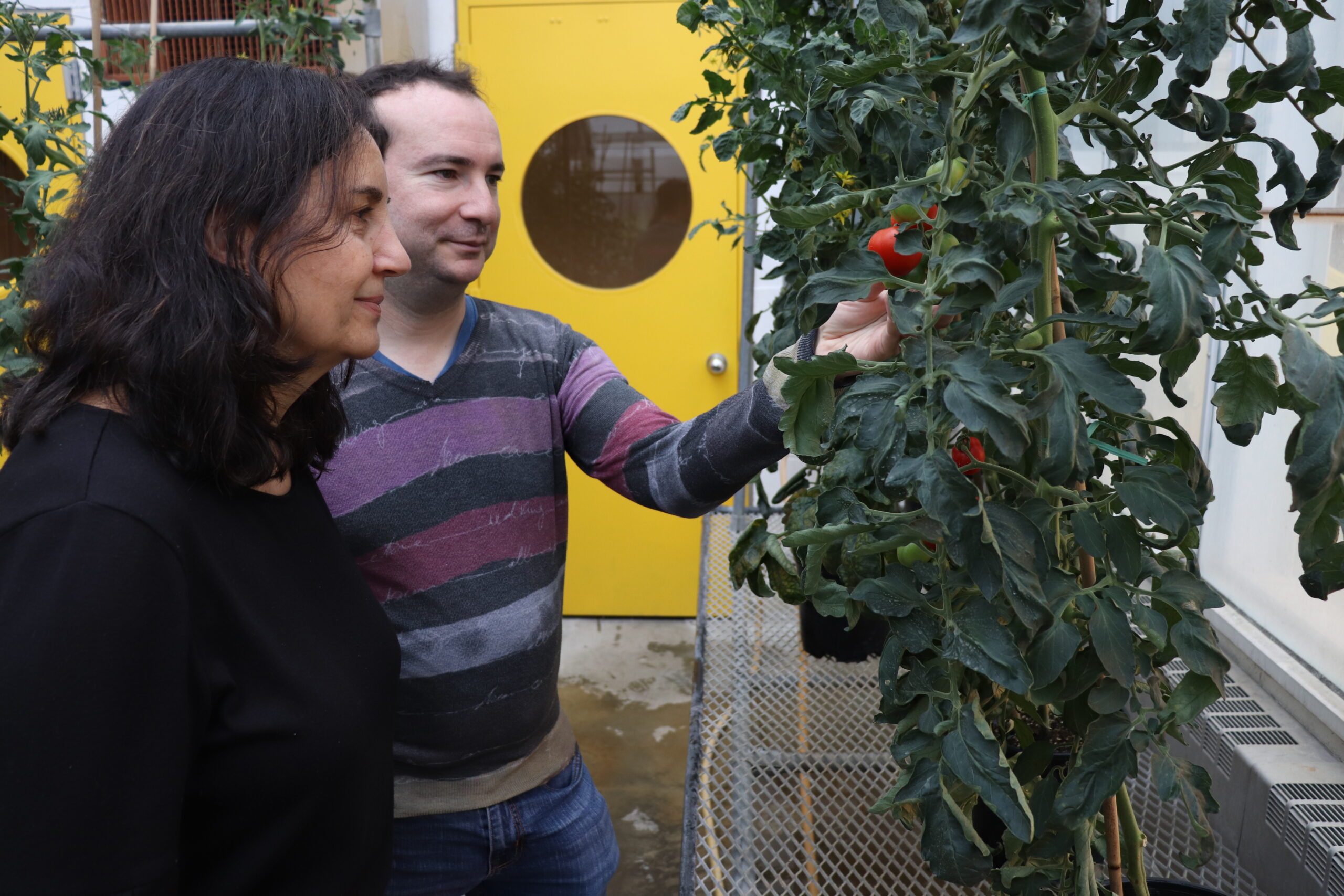
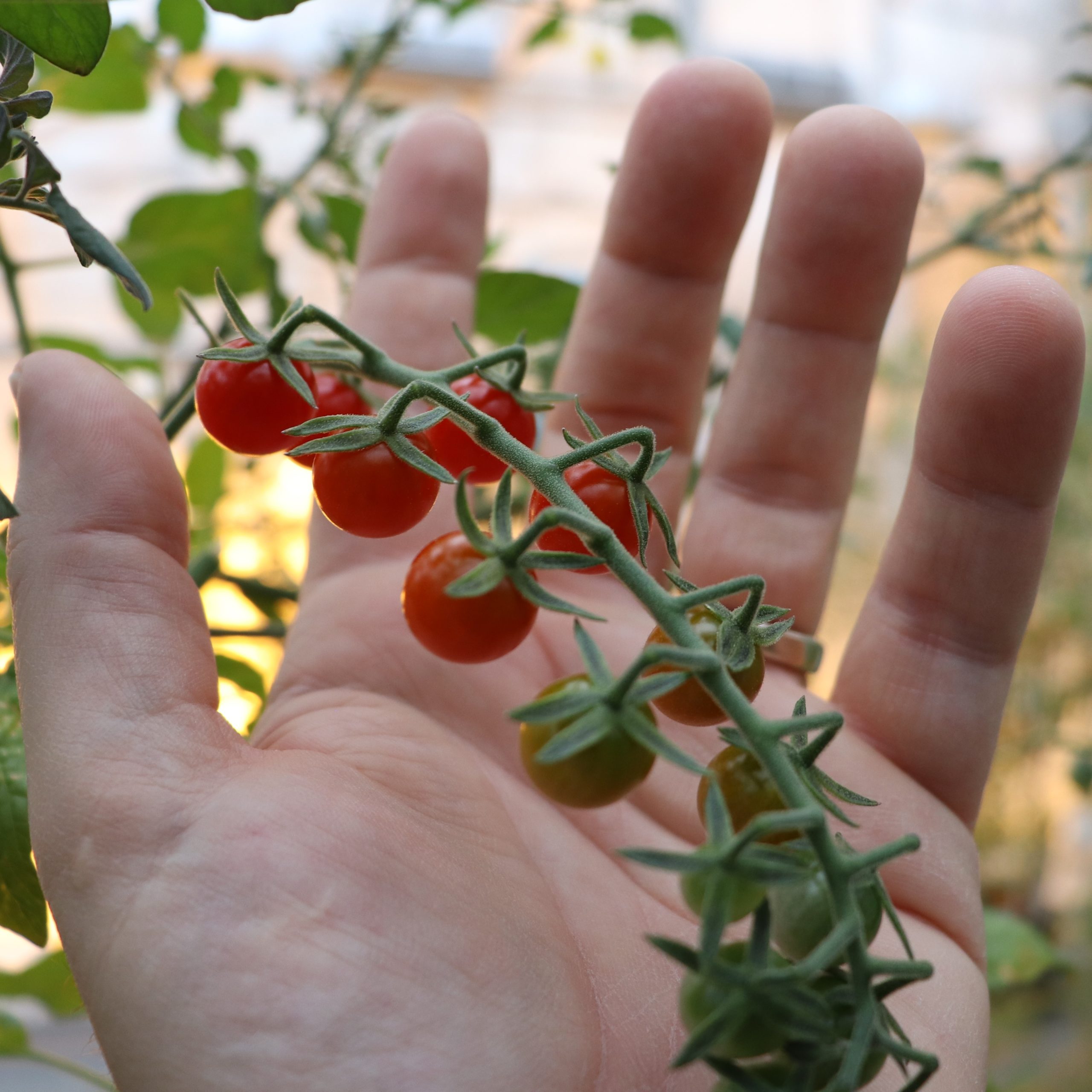
Our research program is focused on elucidating key questions related to auxin synthesis, translocation and the nature of auxin-regulated signaling networks during fruit development, using tomato as a model system.
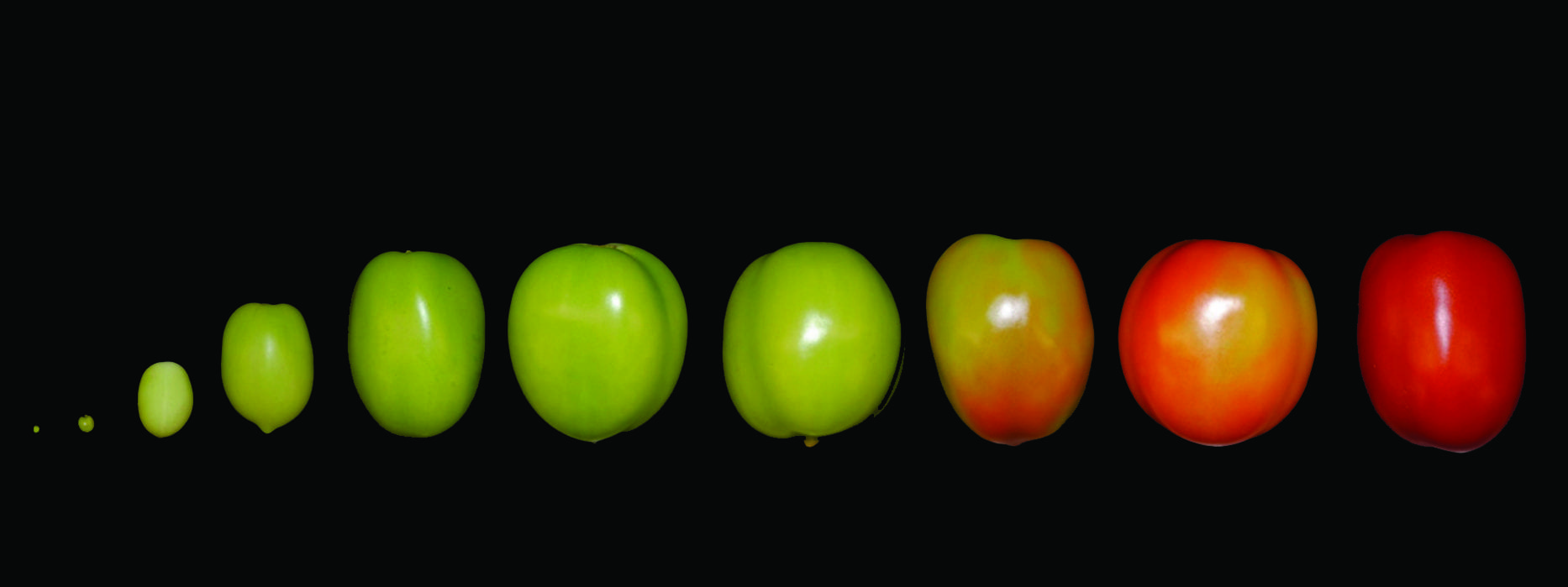
During fruit set, the growth of an otherwise static ovary is stimulated after successful pollination and fertilization. After fertilization, tomato fruit growth is due primarily to cell division and later fruit growth continues mostly by cell expansion. At the end of the cell expansion period, the fruit has reached its final size and will start to ripen.
Auxin homeostasis during tomato fruit growth and development
Despite major advances made in recent years in many aspects of auxin metabolism, transport and signaling in vegetative tissues, the information about the nature and importance of these processes in fruit development and ripening of crop fruit species is very scarce. Moreover a recurring theme that emerges from all these studies is the lack of knowledge about the sources of auxin in fruit tissues, its biosynthetic pathway(s) and how auxin becomes distributed to fruit target tissues. Our research goal is to better understand the mechanisms by which auxin is produced and transported in tomato fruit and how these mechanisms are regulated to mediate cell and tissue specific growth and differentiation.
Analysis of auxin levels or activity in different tomato tissues have revealed a dynamic pattern of tissue specific auxin accumulation throughout fruit development likely to be regulated by components of the auxin polar transport pathway. Critical components of auxin transport systems are the PIN and AUX/LAX protein families, which control cellular auxin efflux and influx respectively. Our studies have provided a transcriptional map for the PIN and AUX/LAX gene families of auxin transport facilitators in the tomato fruit, an important first step towards unraveling the complex network controlling auxin transport routes during fruit set and growth. Multiple PIN and AUX/LAX genes show both overlapping, and tissue-specific patterns of expression suggesting that the coordinated action of PIN and AUX proteins is required for establishing the adequate auxin pools and gradients controlling growth and differentiation in fruit tissues. We also seek to elucidate the mechanisms of IAA biosynthesis in tomato fruit and we are focusing on the tomato orthologs of the tryptophan aminotransferase of Arabidopsis (TAA1) which converts tryptophan into the IAA precursor indole-pyruvic acid and is a key enzyme contributing to IAA production in vivo.
The hypothesis underlying our research is that a tightly regulated spatial and temporal control of auxin levels during tomato fruit development is necessary to activate ovary growth upon fertilization and to coordinate cell expansion and differentiation during exponential fruit growth. We are testing this hypothesis by manipulating the gene expression of specific auxin transporters and auxin biosynthetic genes using fruit-specific promoters and analyzing the effect on fruit development and the dynamics of auxin distribution.
Cell-specific analysis of the tomato fruit transcriptome for the discovery of genes and networks regulating fruit development
One of the first objectives of this research, funded by the NSF Plant Genome Program, is to generate a comprehensive assessment of the cell specific transcript landscape of the developing tomato fruit using Laser Capture Microdissection (LCM) coupled with mRNA profiling by the Illumina platform.
We are mining the tissue-specific transcript datasets for genes associated with hormone signaling, synthesis and transport and with cell wall biosynthesis and modification processes. This non-targeted approach has the potential to dramatically increase the discovery of rare and cell-type specific transcripts and will help identify regulatory hormonal networks controlling auxin homeostasis, as well as new/novel components in the auxin biosynthetic, transport and response pathways. We are using this information to build a model integrating hormone regulated cell expansion and tissue growth and to identify pathways potentially critical to fruit set and growth.
-
Finding genes to help fruit adapt to droughts
As climate change is expected to lead to more frequent periods of drought, researchers are increasingly working to make discoveries that can help plants adapt to prolonged water stress. Researchers […] Read more » -
Tomato’s Wild Ancestor Is a Genomic Reservoir for Plant Breeders
Thousands of years ago, people in the region now known as South America began domesticating Solanum pimpinellifolium, a weedy plant with small, intensely flavored fruit. Over time, the plant evolved […] Read more » -
New ‘Tomato Expression Atlas’ dives deep into the fruit’s flesh
Researchers at BTI, Cornell and USDA published a spatiotemporal map of gene expression across all tissues and developmental stages of the tomato fruit – the genetic information underlying how a fruit changes from inside to out as it ripens. Their data is available in the new Tomato Expression Atlas (TEA). Read more »
-
SolGenomics Meeting Has Newest Advances in Nightshades
Many BTI researchers will present their latest research at the 13th annual SolGenomics Conference, Sept. 12-16 in Davis, California. Read more » -
And the Winners Are…
BTI announces the winning proposals submitted to the Triad Foundation’s Plants and Human Health grant program. Read more » -
From Flower to Fruit: Study Reveals Details of Tomato Formation
BTI Researchers pinpointed which genes are important at different stages of tomato fruit development by monitoring how gene expression changed in the first four days after a flower becomes pollinated. Read more »
Investigating the molecular mechanisms underlying fruit set and development
Fruit development is a crucial process in the sexual reproduction of flowering plants and of critical importance for seed dispersal, plant fitness and agricultural yield. Fruit are complex organs which arise from the coordinated growth and development of floral tissues following pollination. Research in the Catala lab focuses on the molecular regulation of fruit formation and early development using tomato as a model system. We use molecular and genetic techniques to investigate the complex interplay of gene expression changes, signaling events, and hormonal activity, controlling fruit development. The lab also studies the effect of drought stress, an increasing problem in crop production, on tomato fruit set and growth. We are taking advantage of the genetic diversity of wild tomato species, to examine the molecular basis of adaptations to water stress as well as of other fruit quality traits.
Internship Program | Projects & Faculty | Apply for an Internship