News
New ‘Tomato Expression Atlas’ dives deep into the fruit’s flesh
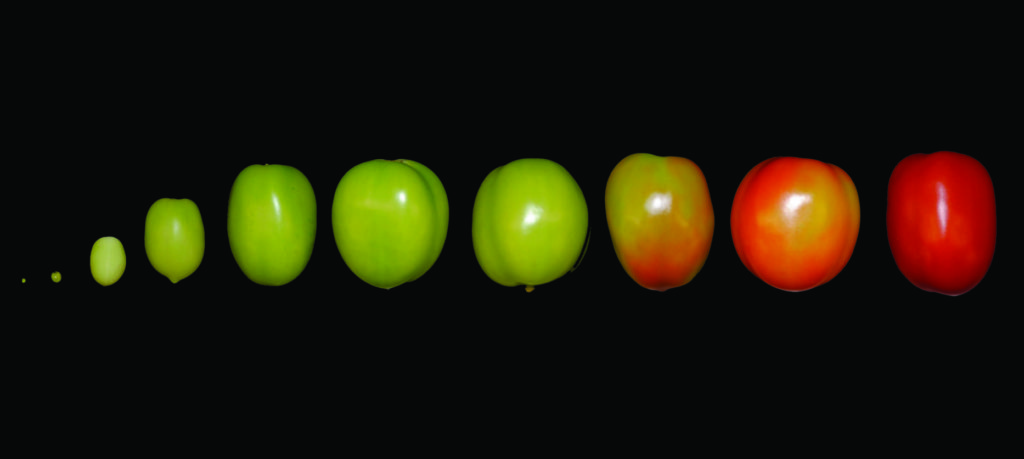

Tomato fruit at the different developmental stages used for this study. Credit: The Tomato Expression Atlas.
From fried green tomatoes to pizza pie, the world savors the tomato across many stages of ripeness, each with its unique qualities. How a fruit ripens has long been an important question for breeders, and the subject of an extensive and fruitful collaboration involving researchers at Boyce Thompson Institute (BTI), Cornell University and the U.S. Department of Agriculture (USDA).
Published online in Nature Communications Jan. 25, the researchers present a spatiotemporal map of gene expression across all tissues and developmental stages of the tomato fruit – the genetic information underlying how a fruit changes from inside to out as it ripens. Their data is available online in the new Tomato Expression Atlas (TEA).
“The collaboration involved a number of research groups, all of whom study tomato fruit biology and have been working together for many years,” said Jocelyn Rose, Cornell professor and lead investigator for the project.
With a $55 billion annual market value, tomato is an important subject for understanding the genetic basis of commercially important traits, such as size, color, flavor, and nutritional content. Furthermore, the tomato serves as a model fruit to understand ripening in a way that can be translated to other, more difficult-to-study fruits.
Most previous research has focused on the tomato fruit as one homogenous tissue, or only the outer fleshy part, but as anyone who has chopped a ripe, runny tomato knows, the fruit is far from uniform. For this project, the researchers analyzed each tissue individually at all stages of development, requiring extreme care, time, and various skills – which is where collaboration was key.
“The project took advantage of the highly synergistic capabilities of the collaborators who combined their respective expertise in physiology, molecular biology, laser capture microdissection, data analysis, and database design,” said James Giovannoni, BTI/USDA scientist.
“Our first objective was to generate a transcriptomic atlas during tomato fruit development with an unprecedented level of spatiotemporal resolution,” said Philippe Nicolas, postdoctoral scientist at BTI. “We needed unbiased sampling that was as representative as possible. For that purpose, we harvested in total more than 400 samples from more than 60 randomly selected individual tomato plants.”
The researchers carefully dissected the tomato tissues by hand and with laser capture microdissection to isolate and sequence RNA, the genetic material that makes each tissue distinct, from individual tissues and even cells. The sequence data was then compiled, parsed, and organized into the TEA, where it can be analyzed to investigate the various biological processes important for fruit development. The rigorous bioinformatics work was done by the Mueller and Fei labs at BTI.
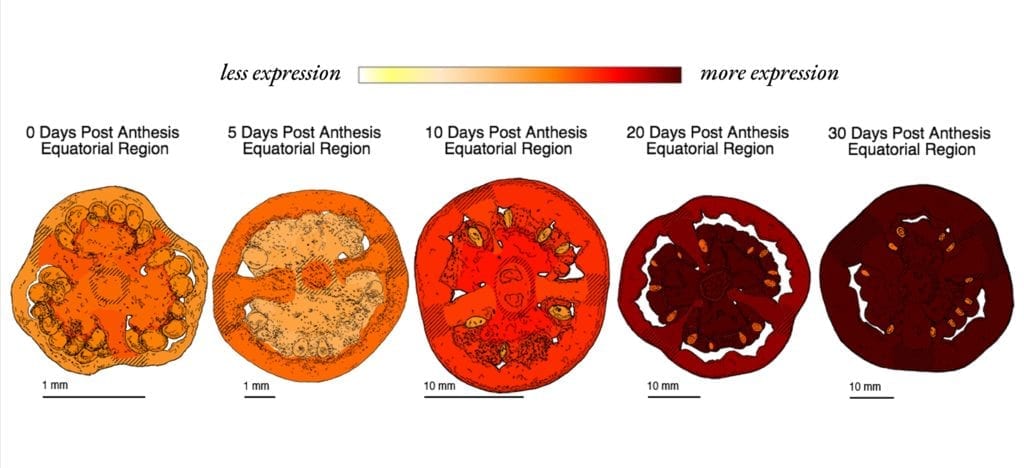
Example of pictographs showing gene expression (Solyc01g102660) over the course of ripening. Credit: The Tomato Expression Atlas.
“The TEA database offers an unprecedented level of interactivity and novel ways to visualize complex, multidimensional expression data,” said BTI scientist Lukas Mueller, referring to the TEA’s graphic interface that allows users to visualize gene expression through heat maps and fruit pictographs.
“These features will enable researchers to easily answer questions that previously were much more tedious to address,” Mueller said.
Just as the collaborating research groups contributed their various expertise to the creation of the TEA, they are each able to use the database to investigate the biological processes important to their own research.
“One of my main research interests is to develop and test models of hormonal regulation of fruit development,” said BTI scientist Carmen Catalá. “We can now use the TEA to generate a comprehensive map of hormonal biosynthesis and signaling pathways encompassing all fruit tissues.”
The Catalá lab used this map to identify two proteins that interact, and likely work together to regulate fruit hormone signaling – one of several TEA-facilitated results in the publication.
“Tomato has been studied for many decades, and many processes have been characterized in considerable detail, but the TEA provides new insights into essentially every process that we’ve examined and gives a high-resolution image of these processes,” Rose said. “It’s rather like holding up a lens to examine a blurred image and having information jump into sharper focus.”
With a clearer image of the biological processes underlying the development of the tomato fruit, researchers can more quickly identify the genetic basis for the many important traits we value in tomatoes and other fruits.
“Our own understanding of the genetic and epigenetic control of ripening and the metabolic pathways that contribute to nutrient quality has been improved,” said Giovannoni. “This greater understanding paves the way for fruit improvement through molecular-assisted breeding.”
This research is supported by grants from the Plant Genome Research Program of the US National Science Foundation, the USDA Agricultural Research Service and the Agriculture and Food Research Initiative of the National Institute of Food and Agriculture.

