News
Scientists Find New Tool for Pathogen to Pillage Plants

Alan Collmer, the Andrew J. and Grace B. Nichols Professor of Plant Pathology and Plant-Microbe Biology at Cornell University and Gregory Martin, the Boyce Schulze Downey Professor at BTI.
Like jewel thieves pulling off a heist, bacteria have a variety of tricks to avoid detection when plundering a plant cell—and researchers continue to learn more details about their highly evolved tactics.
A well-studied protein, AvrPtoB, from the bacterial pathogen Pseudomonas syringae, is known to suppress the plant immune response, but has a previously unknown function, according to researchers from the Boyce Thompson Institute, Cornell University and the USDA-Agricultural Research Service. AvrPtoB prevents the plant from detecting another bacterial protein, HopAD1, which helps the bacterium to reproduce unnoticed. The new study appears in the journal Cell Host & Microbe.
The novel function of AvrPtoB was “like a new trick,” said co-author Alan Collmer, the Andrew J. and Grace B. Nichols Professor of Plant Pathology and Plant-Microbe Biology at Cornell University. “AvrPtoB is like a Swiss Army knife where every time you look, you find another tool.”
P. syringae is a bacterial species that infects a wide variety of plants. In tomatoes, it causes bacterial speck disease and creates brown lesions on leaves and fruits. The bacterium launches an infection by sticking a needle-like tube into the plant cell and injecting proteins called effectors. These proteins attempt to disable plant defense proteins, which the plant uses like security guards to monitor for invaders.
Over evolutionary time, the bacterial and plant populations have engaged in an arms race, where the bacteria evolve new effector proteins to help them sneak in, while the plant acquires new defense proteins to uncover and respond to the attack. The arms race has created an incredibly complex and interconnected system involving more than 30 effector proteins.
Since their first work together in 2000, Collmer and Martin have sought to tease apart the interactions between effectors and plant defense proteins, to understand how and when the plant’s immune system responds. They created a strain of P. syringae that has no effector genes. Then, they put small sets of effector genes back in and tested each strain’s ability to infect leaves of the model plant, Nicotiana benthamiana.
“The early part of this project involved finding all of these effector genes and now we’ve basically knocked all of them out,” said Collmer. Now they must figure out how all the different proteins interact. “It’s a like a giant jigsaw puzzle and we have all of the pieces on the table and the challenge now is how they all fit together.”
Plants have two lines of defense for fighting off bacteria such as P. syringae. Proteins on the cell surface can detect that bacteria are present and mount a weak and transient defense called pattern-triggered immunity (PTI). PTI is like the motion-sensing lights outside a jewelry store that scare off loiterers and stray dogs. But like a skilled jewel thief, the bacterium can outwit PTI by using its effector proteins. One known function of AvrPtoB is to block the actions of the cell surface receptors that trigger PTI.
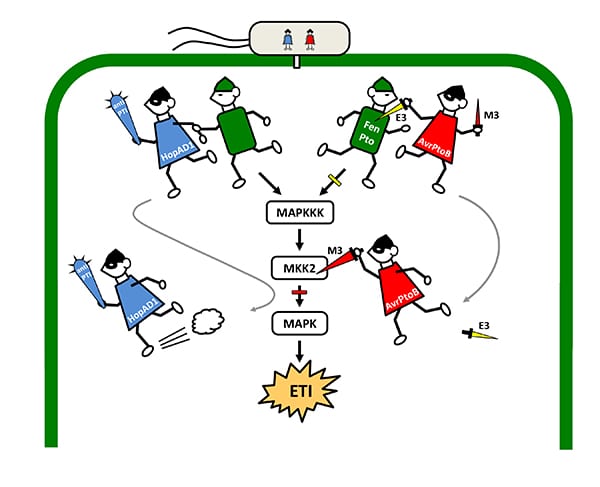
How HopAD1 and AvrPtoB help P. syringae to evade the plant immune system and raid the cell. (Created by Senior Research Associate Magdalen Lindeberg)
In the second line of defense, the plant uses defense proteins inside the cell to detect the bacterial effectors and sound the alarm. This response is called effector-triggered immunity, or ETI, and ultimately results in the plant killing off its own affected cells to prevent the bacteria from replicating and spreading. Previous studies have shown that AvrPtoB blocks two plant defense proteins, Pto and Fen, which helps the bacterium to evade ETI.
“PTI is a little weaker and pathogens can overcome it in many cases. Whereas when an effector is detected inside the plant cell, the plant knows for sure it is being attacked and it activates ETI, a very strong immune response,” said co-author Greg Martin, the Boyce Schulze Downey Professor at BTI. “If a pathogen wants to survive, it needs to figure out how to interfere with ETI.”
By testing various effector proteins, the researchers found that one called HopAD1 could trigger ETI. Additionally, they discovered a previously hidden phenomenon: AvrPtoB can mask the detection HopAD1. It accomplishes this by disabling the MKK2 protein, which the cell needs to trigger ETI.
“We have worked with AvrPtoB since 2002 and thought we understood most aspects of the protein,” said Martin. “It was a surprise to find it still had other tricks up its sleeve.”
The study’s findings underscore why it is difficult for plant breeders to create disease-resistant crops. Even if a specific defense gene enables the plant to detect and stop an infection, the pathogen can evolve a new strategy to overcome the plant’s resistance. A breeder can take 10 years to develop a resistant plant, but often bacteria can overcome that resistance in as little as six years. Now, breeders commonly practice “pyramiding,” where they include multiple defense genes in the plant, to make it harder for the bacterium to evolve an alternate strategy.
“The more we understand the molecular mechanisms involved in the infection process and in plant resistance, the more effective we’re going to be in breeding resistant crops,” said Martin.
For more information on P. syringae, check out the Vegevaders™ board game created by Bryant Adams, Candace Collmer, Alan Collmer, and Magdalen Lindeberg.
Subscribe to BTI's LabNotes Newsletter!
Contact:
Boyce Thompson Institute
533 Tower Rd.
Ithaca, NY 14853
607.254.1234
contact@btiscience.org
Copyright © 2023 | Boyce Thompson Institute | All rights reserved | Privacy Policy | Cookie Policy


