Frank C. Schroeder
Dr. rer. nat.
Professor, Daniel F. Klessig Distinguished Scientist
Development of viable approaches for the systematic identification of the more 100,000 unknown metabolites in animal model systems and the human body, many of which serve essential roles for human health and represent an untapped resource for the development of new drug leads.

Small molecule signaling that regulates host-microbe interactions in animals and plants.
Email: schroeder@cornell.edu
Office Phone: 607-254-4391
Office/Lab: Room 425/414-422
Professor
Boyce Thompson Institute
Professor
Department of Chemistry & Chemical Biology
The College of Arts & Sciences
Cornell University
Graduate Fields:
Chemistry and Chemical Biology
Biochemistry, Molecular & Cell Biology
Host metabolism balances microbial regulation of bile acid signalling. Won TH, Arifuzzaman M, Parkhurst CN, Miranda IC, Zhang B, Hu E, Kashyap S, Letourneau J, Jin WB, Fu Y, Guzior DV; JRI Live Cell Bank; Quinn RA, Guo CJ, David LA, Artis D, Schroeder FC. Nature 2025, 638, 216-224.
Mechanisms of metabolism-coupled protein modifications. Zhang B, Schroeder FC. Nat. Chem. Biol. 2025.
Evolutionarily related host and microbial pathways regulate fat desaturation in C. elegans. Fox BW, Helf MJ, Burkhardt RN, Artyukhin AB, Curtis BJ, Palomino DF, Schroeder AF, Chaturbedi A, Tauffenberger A, Wrobel CJJ, Zhang YK, Lee SS, Schroeder FC. Nat Commun 15, 1520 2024.
Sex-specificity of the C. elegans metabolome. Burkhardt RN, Artyukhin AB, Aprison EZ, Curtis BJ, Fox BW, Ludewig AH, Palomino DF, Luo J, Chaturbedi A, Panda O, Wrobel CJJ, Baumann V, Portman DS, Lee SS, Ruvinsky I, Schroeder FC. Nat Commun. 2023 Jan 19;14(1):320.
Parallel pathways for serotonin biosynthesis and metabolism in C. elegans. Yu J, Vogt MC, Fox BW, Wrobel CJJ, Fajardo Palomino D, Curtis BJ, Zhang B, Le HH, Tauffenberger A, Hobert O, Schroeder FC. Nat Chem Biol. 2023 Feb;19(2):141-150.
- US Patent: 12382888B2
- US Patent: 12376588B2
- US Patent: 12,180,246
- US Patent: 11,464,810
- European Patent: EP3503904B1
- US Patent: 11,077,151
- US Patent: 10,136,595 / 11,019,776 / 11,849,688
- European Patent: EP2,967,052
- Chinese Patent: CN105263324
- Canadian Patent: CA2907470A1
- US Patent: 9,868,754 / 10,183,963 / 10,479,813 / 11,673,908
- US Patent: 9,534,008
- Chinese Patent: CN103841827
- Japanese Patent: JP6109829
- US Patent: 9,487,551
- US Patent: 9,445,596
Research Overview
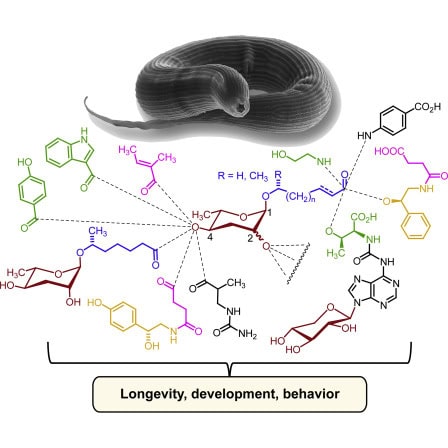
Our research is dedicated to facilitating systematic structural and functional annotation of small biogenic molecules – the metabolome – in model organisms, integrating expertise in molecular biology, bioinformatics, and analytical chemistry. Structurally diverse but yet unidentified metabolites play central roles as information carriers in most biological processes and represent an increasingly recognized source for new drug leads, e.g., the treatment of metabolic disease and cancer. Moreover, detailed knowledge of small-molecule structures, their biosynthetic pathways, and their interactions with other biomolecules is essential for advancing our understanding of endocrine and exocrine signaling pathways.
Combining NMR-spectroscopic and MS-based comparative metabolomics with phenotypic screens and genetic approaches, we have engaged in a comprehensive effort to characterize structures and functions of the metabolomes of animal model organisms, focusing on metabolites that control development, lipid metabolism, and interactions with the microbiota. Our research has led to the identification of several 100 new metabolites that function as signaling molecules in C. elegans regulating virtually all aspects of physiology. This large diversity of metabolites is derived from the modular assembly of building blocks from conserved metabolic pathways, presenting a new paradigm of small molecule biosynthesis in metazoans.
Encouraged by the initial success of our methods in the model system C. elegans, we engaged in a major effort toward a systematic characterization of mammalian metabolomes, with a focus on changes in response to alteration of the composition of the gut microbiome. These studies indicate that mammals employ similarly extensive small molecule-based signaling networks that rely to a significant extent on not yet annotated metabolites.
Many of the newly identified molecules, including a new class of bile acid conjugates, have entirely unexpected structures or biological activities, and the elucidation of their roles in signaling pathways regulating metabolic homeostasis has become a major focus of my lab. This includes elucidation of the biosynthetic pathways that regulate abundance of the identified signaling molecules, identification of corresponding receptors, and their associated physiological phenotypes.
To accelerate detection and functional characterization of new metabolites my lab has developed a suite of software tools (“Metaboseek”) for the research community that facilitate comparative metabolomic analysis of high-resolution MS data to detect metabolites associated with specific mutations, developmental stages, or environmental conditions. As a necessary component of the chemical biology-related aspects of our research, my lab has extensive experience with syntheses of newly identified metabolites (for phenotypic assays) and related compounds, e.g. probes to identify receptors or interrogate biosynthetic pathways.
Please see our Group Website for recent news and publications, research updates, and teaching.
Lab Members

Arnaud Tauffenberger
Research Associate
Arnaud investigates the metabolic processes underlying ageing and neurodegeneration using C. elegans.
Name pronunciation: https://www.youtube.com/watch?v=ZTkfUb18d2s

Bennett Fox
Research Associate

Goncalo Gouveia
Research Associate
Gonçalo is an analytical chemist by training and is developing computational tools to improve metabolite annotation and compound discovery as well as methods to improve reproducibility of metabolomics with holistic strategies from experimental design to data analysis.

Tyler Schwertfeger
Graduate Student
Tyler develops synthetic and mass spectrometry-based approaches for characterizing new RNA modifications in C. elegans.

Allen Schroeder
Graduate Student
Allen is employing synthesis and metabolomics to probe metabolic pathways involved in neurotransmitter biosynthesis and fatty acid degradation.

Elise N. Garner
Graduate Student
Elise is identifying metabolic pathways connected to mTORC2 using untargeted metabolomics.

Samuel Kemiji
Graduate Student
Thesis advisor: Sijin Lin

Marissa A. Fontaine
Graduate Student
Marissa's research aims to explore the complex host-microbiome interplay by employing a combination of targeted and untargeted comparative metabolomics as well as proteomics in mouse models.

Olivia Duncan
Graduate Student
Olivia is investigating the role of the gut microbiome in directing host fat metabolism.

Yu-Heng Hsieh
Graduate Student
Yu-Heng is using metabolomics and proteomics to study plant-microbe symbiosis.

Yuxin Xie
Graduate Student
Yuxin is employing a combination of analytical tools and total synthesis to understand nucleotide catabolism in C. elegans, as well as RNA modifications in the closely related P. pacificus.

Tomer Markovich
Graduate Student
Tomer is interested in the small molecules involved in stem cell signalling, specifically in the context of extracellular vesicles.

Ali Turk
Graduate Student
Ali is investigating mitochondrial metabolite transport and the role of the electron transport chain in regulating catabolism in C. elegans .

Annie Liu
Graduate Student
Annie is interested in investigating the nucleus via proteomics and metabolomics.

Chiara Giraldo

Haofan Wang
Graduate Student

Leah Kunkee
Graduate Student

Laura D. Galindo
Research Assistant

Ryan T. Moradei
Undergraduate Student

Daniel N. Salter
Undergraduate Student

Sky Chen
Undergraduate Student

Alex S. Cerreno
Undergraduate Student
In the News

How Bacteria Use Sneaky Chemistry to Disable Plant Defenses
In the microscopic battlefield of plant-microbe interactions, plants are constantly fighting off invading bacteria. New research reveals just how clever these bacterial invaders can be. Plants, like humans, have evolved...
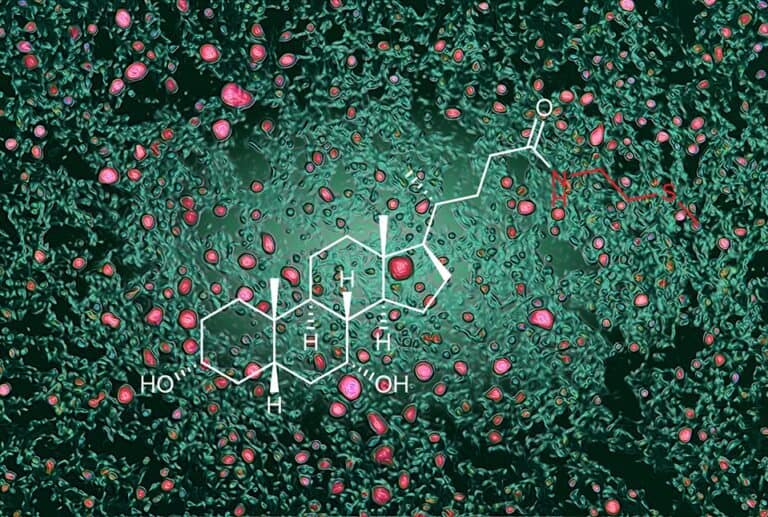
Diet, Microbes and Fat: A New Pathway Controlling Levels of Body Fat and Cholesterol
Beneficial gut microbes and the body work together to fine-tune fat metabolism and cholesterol levels, according to a new preclinical study by investigators from Weill Cornell Medicine and the Boyce Thompson Institute....
Synopsis of paper: Evolutionarily related host and microbial pathways regulate fat desaturation in C. elegans
Written by Bennett Fox In our paper, recently published in Nature Communications, we (my colleagues at the Boyce Thompson Institute and Cornell University) demonstrated that a specific type of bacteria-derived fatty...
Mounting evidence suggests that the secret to understanding human health and combating metabolic diseases lies hidden within the microscopic world of our gut bacteria. Recent research by scientists at the Boyce...
Tiny Worm, Giant Leap: Discovery of Highly Specific Fatty Acid Attachment to Proteins
In a world where the intricacies of molecular biology often seem as vast and mysterious as the cosmos, a new groundbreaking study delves into the microscopic universe of proteins, unveiling...
Deciphering Molecular Mysteries: New Insights into Metabolites That Control Aging and Disease
In a significant advancement in the field of biochemistry, scientists at BTI and Cornell University have uncovered new insights into a family of metabolites, acylspermidines, that could change how we understand...