News
A New Method for Studying Plant-Germ Warfare
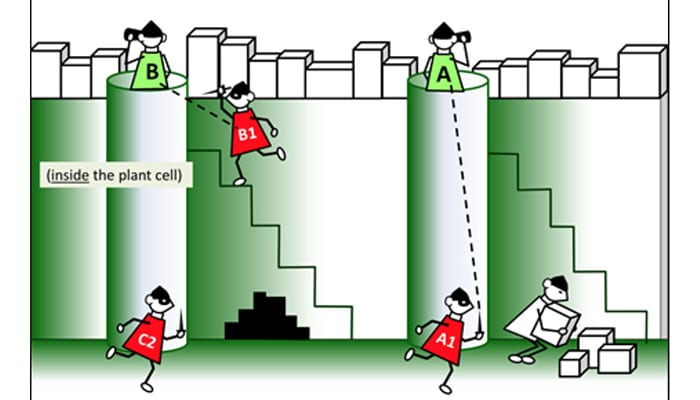
Invading bacteria hijack the plant cell machinery to decorate their attack proteins (in red) with fatty acids, so that plant will transport them to the greasy cell membrane. Once they are at the surface, the attack proteins attempt to disable the plant’s bacterial sensors (in green). (Image from Vegevaders, courtesy of Magdalen Lindeberg of Cornell University)
In the evolutionary arms race between plants and invading bacteria, the greasy plasma membrane at the surface of the plant cell is a veritable hotbed of sneaky plant-pathogen interactions.
In the search to untangle molecular tactics used in this arms race, researchers in the lab of Gregory Martin at the Boyce Thompson Institute (BTI) have developed a technology to determine if a protein is with fatty acids in the plant cell—a modification that typically traffics the protein to the plasma membrane. Plant cells use these fatty acid modifications to install sensors for bacterial infection at their membrane, but bacteria hijack the plant’s fatty acid modification machinery to traffic their own proteins, called effectors, to this critical plant-pathogen battleground.
The researchers applied their technique to two different sensors from the plant and a bacterial effector, all of which are known to be fatty acid modified, to provide a proof of concept. The method, is described in an article in the Sept. 2 issue of Plant Methods, and may be useful for studying a variety of plant proteins and pathogen effectors.
“The membrane of the plant cell is greasy,” explained co-author Patrick Boyle, formerly a postdoctoral scientist in the Martin lab and now a researcher at Monsanto Co. in the Emerging Leaders in Science program. “When a disease-causing bacterium is trying to invade the plant, it injects these effector proteins behind enemy lines, inside the plant cell. These proteins are molecular saboteurs. They go in and cut all the surveillance machinery wires so the plant cell can’t sound the alarm and turn on its defenses.” In some cases, effectors get to the bacteria-detecting sensors in the plant plasma membrane by greasing themselves up with fatty acids from the plant cell.
The method detects two major forms of fatty acid modification, N-myristoylation and S-acylation, through click chemistry, which uses chemical reactions that work in biological systems and can link a biomolecule to a reporter molecule. The researchers simply inject a plant with a slightly altered fatty acid and allow the plant’s own inherent metabolic mechanisms to incorporate it into the protein of interest. They then purify the complex from the plant and “click” on the reporter molecule that makes it light up for detection. This approach has been used previously to detect protein fatty acid modifications in various systems but the Martin lab adapted the technology to study such modification in the context of plants.
To show that the system works, the researchers used the technique to detect the fatty acid modification of a bacteria-detecting receptor, FLS2, in plant cells. They also applied the technology to show modification of both the AvrPto effector from the bacterium Pseudomonas syringae pv. tomato, and Pto, the plant immunity protein that perceives this effector, in leaves of the model tobacco plant Nicotiana benthamiana. In the future, however, scientists could potentially leverage this technology to capture and identify unknown proteins labeled through N-myristoylation and S-acylation, from all the proteins in the plant cell.
Some of the traditional methods for detecting fatty acid-modified proteins require the use of radioactive compounds and can take months of incubating to yield a result. “This technique is radiolabel-free, reduces detection time to a few days and can complement other approaches for studying the fatty acid modification of proteins in plants,” said Boyle.
The system offers a simple and rapid tool for teasing apart the membrane-level measures and countermeasures used by plants and infectious bacteria during an attack.
“It will help shine light on some of the greasier tactics used in this ever-escalating molecular arms race between the plant and the pathogen,” said Boyle.
Additional researchers on the paper include co-first author and former graduate student Simon Schwizer, research associate Sarah Hind, graduate student Christine Kraus, former research assistant Susana De la Torre Diaz, and Bin He, currently at Guiyang Medical University.
The study received support from the U.S. Department of Agriculture—National Institute of Food and Agriculture, the National Science Foundation, the Human Frontier Science Program and the Triad Foundation.
Subscribe to BTI's LabNotes Newsletter!
Contact:
Boyce Thompson Institute
533 Tower Rd.
Ithaca, NY 14853
607.254.1234
contact@btiscience.org
Copyright © 2023 | Boyce Thompson Institute | All rights reserved | Privacy Policy | Cookie Policy


